

收稿日期: 2018-05-08
网络出版日期: 2018-12-23
基金资助
国家自然科学基金资助项目(81430083)
Experimental methods for characterizing protein-protein interactions
Received date: 2018-05-08
Online published: 2018-12-23
生物体内的蛋白质分子常常通过与其他蛋白质分子发生相互作用来发挥其生物功能. 因此, 研究蛋白质-蛋白质相互作用 (protein-protein interaction, PPI) 对于阐明蛋白质分子的生物功能以及分子作用机理具有重要的意义. 主要介绍基于生物物理和生物化学原理的检测蛋白质-蛋白质相互作用的实验研究方法, 并对发展趋势做出展望.
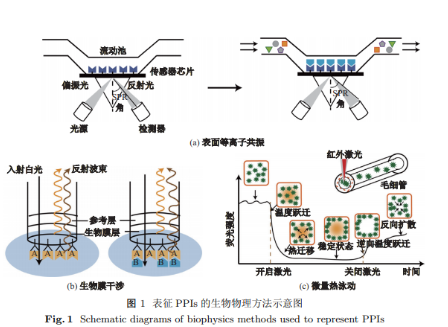
关键词: 蛋白质-蛋白质相互作用; 生物物理方法; 生物化学方法
孔文娜, 周宓, 林海霞, 王任小 . 蛋白质-蛋白质相互作用的实验检测方法[J]. 上海大学学报(自然科学版), 2021 , 27(1) : 106 -116 . DOI: 10.12066/j.issn.1007-2861.2065
Protein molecules usually interact with other partner protein molecules to execute their biological functions. Hence, the characterisation of protein-protein interac- tions (PPIs) plays a significant role inelucidating the molecular mechanism of proteins. With the progress of proteomics comes the development of a variety of experimental methods for characterising protein-protein interactions. This article mainly reviews recently developed biophysical- or biochemical-basedexperimental methods for characterising protein-protein interactions.

| [1] | Berggard T, Linse S, James P. Method for the detection and analysis of protein-protein interactions[J]. Proteomics, 2007,7:2833-2842. |
| [2] | Pandey A, Mann M. Proteomics to study genes and genomes[J]. Nature, 2000,405(6788):837-846. |
| [3] | Villoutreix B O, Kuenemann M A, Poyet J L, et al. Drug-like protein-protein interaction modulators: challenges and opportunities for drug discovery and chemical biology[J]. Molecular Informatics, 2014,33:414-437. |
| [4] | Fernández-Recio J. Prediction of protein binding sites and hot spots[J]. WIREs Computational Molecular Science, 2011,1(5):680-698. |
| [5] | Ruffner H, Baver A, Bovwmester T. Human protein-protein interaction networks and the value for drug discovery[J]. Drug Discovery Today, 2007,12(17/18):709-716. |
| [6] | Shimada S, Shinzawa-Itoh K, Baba J, et al. Complex structure of cytochrome c-cytochrome c oxidase reveals a novel protein-protein interaction mode[J]. The EMBO Journal, 2017,36(3):291-300. |
| [7] | Patil S P, Fink M A, Enley E S, et al. Identification of small-molecule inhibitors of PD-1/PD-L1 protein-protein interaction[J]. Chemistry Select, 2018,3:2185-2189. |
| [8] | Thakur S, Dhiman M, Tell G, et al. A review on protein-protein interaction network of APE1/Ref-1 and its associated biological functions[J]. Cell Biochemistry and Function, 2015,33:101-112. |
| [9] | Lemos A, Le?o M, Soares J, et al. Medicinal chemistry strategies to disrupt the p53-MDM2/MDMX interaction[J]. Medicinal Research Reviews, 2016,36(5):789-844. |
| [10] | Zhou M, Li Q, Wang R X. Current experimental methods for characterizing protein-protein interactions[J]. ChemMedChem, 2016,11:738-756. |
| [11] | Rouck J E, Krapf J E, Roy J, et al. Recent advances in nanodic technology for membrane protein studies (2012---2017)[J]. FEBS Letters, 2017,591(14):2057-2088. |
| [12] | Wu L, Tang H L, Hu S Q, et al. Sensitive and simultaneous surface plasmon resonance detection of free and p53-bound MDM2 proteins from human sarcomas[J]. Analyst, 2018,143(9):2029-2034. |
| [13] | Boozer C, Kim G, Cong S X, et al. Looking towards label-free biomolecular interaction analysis in a high-throughput format: a review of new surface plasmon resonance technologies[J]. Current Opinion Biotechnology, 2006,17:400-405. |
| [14] | Concepcion J, Witte K, Wartchow C, et al. Label-free detection of biomolecular interactions using BioLayer interferometry for kinetic characterization[J]. Combinatorial Chemistry & High Throughput Screening, 2009,12:791-800. |
| [15] | Sultana A, Lee J E. Measuring protein-protein and protein-nucleic acid interactions by biolayer interferometry [J]. Current Protocols in Protein Science, 2015, 79: 19.25.1-19.25.26. |
| [16] | Pogoutse A K, Lai C L, Ostan N, et al. A method for measuring binding constants using unpurified in vivo biotinylated ligands[J]. Analytical Biochemistry, 2016,501:35-43. |
| [17] | Wienken C J, Baaske P, Rothbauer U, et al. Protein-binding assays in biological liquids using microscale thermophoresis[J]. Nature Communications, 2010,1:100. |
| [18] | Seidel S A, Dijkman P M, Lea W A, et al. Microscale thermophoresis quantifies biomolecular interactions under previously challenging conditions[J]. Methods, 2013,59(3):301-315. |
| [19] | Mao Y X, Yu L L, Yang R, et al. A novel method for the study of molecular interaction by using microscale thermophoresis[J]. Talanta, 2015,132:894-901. |
| [20] | Jerabek-Willemsen M, Wienken C J, Braun D, et al. Molecular interaction studies using microscale thermophoresis[J]. Assay and Drug Development Technologies, 2011,9(4):342-353. |
| [21] | Arbel N, Ben-Hail D, Shoshan-Barmatz V. Mediation of the antiapoptotic activity of Bcl-xL protein upon interaction with VDAC1 protein[J]. Journal of Biological Chemistry, 2012,287(27):23152-23161. |
| [22] | Lin C C, Melo F A, Ghosh R, et al. Inhibition of basal FGF receptor signaling by dimeric Grb2[J]. Cell, 2012,149(7):1514-1524. |
| [23] | Bhogaraju S, Cajanek L, Fort C, et al. Molecular basis of tubulin transport within the cilium by IFT74 and IFT81[J]. Science, 2013,341:1009-1012. |
| [24] | Jerabek-Willemsen M, André T, Wanner R, et al. Microscale thermophoresis: interaction analysis and beyond[J]. Journal of Molecular Structure, 2014,1077:101-113. |
| [25] | Deng S H, Chen J F, Gao Z S, et al. Effects of donor and acceptor's fluorescence lifetimes on the method of applying F?rster resonance energy transfer in STED microscopy[J]. Journal of Microscopy, 2018,269(1):59-65. |
| [26] | Leavesley S J, Britain A L, Cichon L K, et al. Assessing FRET using spectral techniques[J]. Cytometry Part A, 2013,83A:898-912. |
| [27] | Vu L T, Nguyen T T K, Alam S, et al. Changing blue fluorescent protein to green fluorescent protein using chemical RNA editing as a novel strategy in genetic restoration[J]. Chemical Biology & Drug Design, 2015,86:1242-1252. |
| [28] | Liu R, Hu X J, Ding Y. Rational design of a pH-insensitive cyan fluorescent protein CyPet2 based on the CyPet crystal structure[J]. FEBS Letters, 2017,591:1761-1769. |
| [29] | Otsubo R, Kim M, Lee J, et al. Midori-ishi Cyan/monomeric Kusabira-orange-based fluorescence resonance energy transfer assay for characterization of various E3 ligases[J]. Genes to Cells, 2016,21:608-623. |
| [30] | Aker J, Hesselink R, Engel R, et al. In Vivo hexamerization and characterization of the arabidopsis AAA ATPase CDC$_{4}$8A complex using forster resonance energy transfer-fluorescence lifetime imaging microscopy and fluorescence correlation spectroscopy[J]. Plant Physiology, 2007,145:339-350. |
| [31] | Song T, Madahar V, Liao J Y. Development of FRET assay into quantitative and high-throughput screening technology platforms for protein-protein interactions[J]. Ann Biomed Eng, 2011,39:1224-1234. |
| [32] | Kost L J, Mootz H D. A FRET Sensor to Monitor Bivalent SUMO--SIM Interactions in SUMO Chain Binding[J]. ChemBioChem, 2018,19:177-184. |
| [33] | Mahajan P N, Linder K, Berry G, et al. Bcl-2 and Bax interactions in mitochondria probed with green fluorescent protein and fluorescence resonance energy transfer[J]. Nature Biotechnology, 1998,16:547-552. |
| [34] | Yegorova S, Chavaroche A E, Rodriguez M C, et al. Development of an AlphaScreen assay for discovery of inhibitors of low-affinity glycan-lectin interactions[J]. Analytical Biochemistry, 2013,439:121-131. |
| [35] | Zhang M, Wisniewski J A, Ji H T. AlphaScreen selectivity assay for $\beta $-catenin/B-cell lymphoma 9 inhibitors[J]. Analytical Biochemistry, 2015,469:43-53. |
| [36] | Hill S J, Williams C, May L T. Insights into GPCR pharmacology from the measurement of changes in intracellular cyclic AMP: advantages and pitfalls of differing methodologies[J]. British Journal of Pharmacology, 2010,161(6):1266-1275. |
| [37] | Moad H E, Pioszak A A. Selective CGRP and adrenomedullin peptide binding by tethered RAMP-calcitonin receptor-like receptor extracellular domain fusion proteins[J]. Protein Science, 2013,22:1775-1785. |
| [38] | Harrison A T, Kriel F H, Papathanasopoulos M A, et al. The evalution of statins as potential inhibitors of the LEDGF/p75-HIV-1 integrase interaction[J]. Chemical Biology & Drug Design, 2015,85:290-295. |
| [39] | Ogita N, Okushima Y, Tokizawa M, et al. Identifying the target genes of SUPPRESSOR OF GAMMARESPONSE 1, a master transcription factor controlling DNA damage response in Arabidopsis[J]. The Plant Journal, 2018,94:439-453. |
| [40] | Sierecki E, Stevers L M, Giles N, et al. Rapid mapping of interactions between human SNX-BAR proteins measured in vitro by AlphaScreen and single-molecule spectroscopy[J]. Molecular Cellular Proteomics, 2014,13:2233-2245. |
| [41] | Wagstaff K M, Rawlinson S M, Hearps A C, et al. An AlphaScreen-based assay for high-throughput screening for specific inhibitors of nuclear import[J]. Journal of Biomolecular Screening, 2011,16(2):192-200. |
| [42] | Montse M, Salvador V, Francesc X. Protein complementation assay: approaches for the in vivo analysis of protein interactions[J]. FEBS Letters, 2009,583:1684-1691. |
| [43] | Shibasaki S, Sakata K, Ishii J, et al. Development of a yeast protein fragment complementation assay (PCA) system using dihydrofolate reductase (DHFR) with specific additives[J]. Applied Microbiology Biotechnology, 2008,80(4):735-743. |
| [44] | Nord O, Gustrin A, Nygren P A. Fluorescent of b-lactamase activity in living Escherichia coli cells via esterase supplementation[J]. FEMS Microbilogy Letters, 2005,242:73-79. |
| [45] | Ozawa T, Nogami S, Sato M, et al. A fluorescent indicator for detecting protein-protein interactions in vivo based on protein splicing[J]. Analytical Chemistry, 2000,72(21):5151-5157. |
| [46] | Kim S B, Sato M, Tao H. Split Gaussia luciferase-based bioluminescence template for tracing protein dynamics in living cell[J]. Analytical Chemistry, 2009,81(1):67-74. |
| [47] | Rochette S, Gagnon-Arsenault I, Diss G, et al. Modulation of the yeast protein interactome in response to DNA damage[J]. Journal of Proteomics, 2014,100:25-36. |
| [48] | Liu T Y, Chou W C, Chen W Y, et al. Detection of membrane protein-protein interaction in planta based on dual-intein-couple tripartite split-GFP association[J]. Plant Journal, 2018,94:426-438. |
| [49] | Luo L, King N P, Yeo J C, et al. Single-step protease cleavage elution for identification of protein-protein interactions from GST pull-down and mass spectrometry[J]. Proteomics, 2014,14:19-23. |
| [50] | Schulte R, Talamas J, Doucet C, et al. Single bead affinity detection (SINBAD) for the analysis of protein-protein interactions[J]. PLoS One, 2008,3:e2061. |
| [51] | Kamanova J, Sun H, Lara-Tejero M, et al. The salmonella effector protein SopA modulates innate immune responses by targeting TRIM E3 ligase family members[J]. PLoS Pathogens, 2016,12(4):e1005552. |
| [52] | Fedosejevs E T, Liu L N C, Abergel M, et al. Coimmunoprecipitation of reversibly glycosylated polypeptide with sucrose synthase from developing castor oilseeeds[J]. FEBS Letters, 2017,591:3872-3880. |
/
| 〈 |
|
〉 |